HOW GENES ARE INHERITED: Cystic fibrosis, Dihybrid Inheritance and when Mendel's laws don't apply.
Cystic fibrosis: an example of monohybrid inheritance. The disease is caused by the mutation of a single allele. A normal allele (C) makes a membrane protein essential for the proper functioning of certain epithelial cells. The mutated allele (c) does not code for a functional protein. Most people have two healthy alleles: their genotype is CC. In the early year 2000, around 1 person in 25 caries a faulty allele in the UK population. These carriers (Cc) are perfectly healthy because they also possess a normal allele, which makes the working protein.source
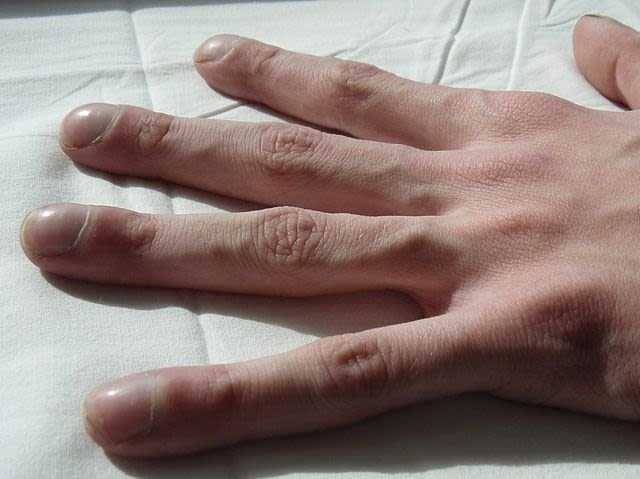
"Clubbing" of the fingers is a classic feature of Cystic Fibrosis. Jerry Nick, M.D, CC BY-SA 3.0
However, in 1 couple in 625 (1 in 25 x 1 in 25) both partners are carriers. There is a one in four chance that any child they have could inherit both faulty alleles and therefore be a cystic fibrosis sufferer. As 1 cystic fibrosis child in 4 is born to 1 Couple in 625, approximately 1 child in every 2500 has this condition.source
DIHYBRID INHERITANCE: HOW TWO GENES ARE PASSED ON
What happens when a tall, purple-flowered pea plant is crossed with a dwarf, white-flowered individual? Do you always get tall plants with purple flowers, or can the features re-combine, producing tall, white-flowered plants, and dwarf, purple-flowered ones? The simple answer is yes. You can get these recombinants, thanks to the process of meiosis.
The inheritance of two separate genes is called dihybrid inheritance. If, as is usual, the two genes are on separate chromosomes, Mendel’s second law applies: each of a pair of contrasted characteristics can be combined with each of another pair. So, an organism with a genotype of AaBb can produce gametes of AB Ab, aB or ab. In other words, one allele from each pair passes into the gamete, and all four combinations are possible. This is due to independent assortment during meiosis.
For an example, let’s return to pea plants. In addition to tall and dwarf. We can consider a gene for seed texture. The allele R for round seeds is dominant to r, the allele for wrinkled seeds.
In the cross between a pure breeding, tall round-seeded plant (TTRR), and a pure breeding dwarf, wrinkled-seeded plant (ttrr). One allele from each pair is passed into the gametes. All of the gametes from the tall round-seeded plant are TR and all those from the dwarf, wrinkled-seeded plant are tr. All of the F1 generation will have the genotype TtRr and are tall with round seeds.
When we cross two TtRr individuals. Things get more complex! Each plant produces four different gametes. Fertilization is random, producing 16 different combinations in the F2 generation. To organize our thoughts we can use a Punnett square.
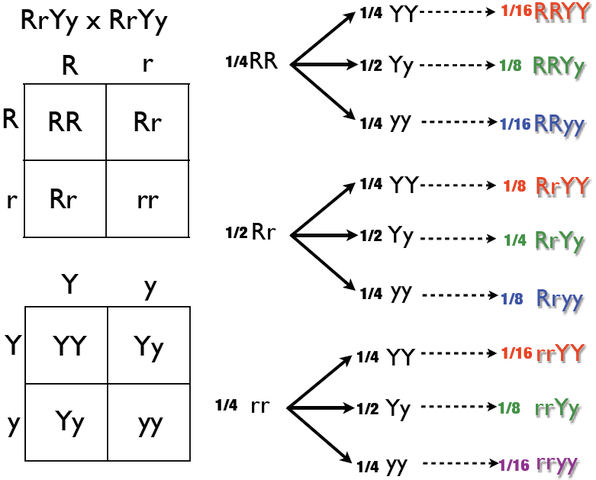
RrYy x RrYy Dihybrid Cross Tree Method using the Punnett square. Tim DeJulio, Public Domain
Dihybrid inheritance questions in examinations
Many examination questions require the candidate to work out the ratios of offspring resulting from a particular cross. If you have not practised these questions they can be very time consuming, leading to stress and silly mistakes.
The example given above (TtRr × TtRr) is the most complex dihybrid situation you can get. This is because both male and female are heterozygous for both alleles, and so produce 4 different gametes each, resulting in 16 different combinations. All other genotypes will produce fewer combinations. If you practise these combinations you can learn to work out the gametes and resulting ratios in your head without resorting to Punnett squares and counting the results.
Consider the cross TtRr × TtRr. The first individual produces the gametes TR and tR in equal amounts. The second one produces Tr and tr in equal amounts. So, there are four possible combinations: the same number as in a monohybrid cross.
You should see that the offspring will be one quarter TTRr. Two quarters TtRr and one quarter ttRr, giving three tall plants with round seeds to one dwarf plant with round seeds. You would not get any dwarf plants with wrinkled seeds. Some exam questions give the ratios of offspring phenotypes and ask you to work backwards to the original genotypes. If you do not get the expected ratios, consider linkage as an explanation below.
WHEN MENDEL’S LAWS DON’T APPLY: LINKAGE AND CROSSING OVER
So far, we have considered the inheritance of genes on different chromosomes that could be separated at meiosis. We say that genes that are on the same chromosome are linked. Such genes cannot obey Mendel’s laws because they do not undergo independent assortment. They are inherited together unless separated by crossing over during prophase 1 of meiosis. In humans, there are over 30 000 genes present on jiust 23 pairs of chromosomes. So, any single gene is linked with a few thousand others because they are all on the same chromosome.
So, to illustrate linkage, let’s consider the sweet pea plant (Lathyrus odoratus – a different species from the edible pea I have been talking about). In this species, the allele for purple flowers is dominant to red flowers, and the allele for elongated pollen grains is dominant to round pollen. If pure breeding plants with purple flowers and elongated pollen grains are bred with plants with red flowers and round pollen grains, the F1 generation all have purple flowers and elongated pollen.
When the F2 are selfed (bred together) you would expect a 9:3:3:1 ratio, but this does not happen. The F2 generation are mainly like the parents (purple and elongated or red and round) but there are a few recombinants (red and elongated or purple and round). This is because the genes for flower colour and pollen shape are linked (present on the same chromosome) and so are inherited together. Any recombinants we get must therefore be due to crossing over.
Consider the set of results shown in the table below. The total number of offspring is 720, of which 59 (31 + 28) are recombinants. Eight per cent of the offspring result from crossing over and so, by convention, we can say that the loci for flower colour and pollen shape are 8 units apart on the chromosome. If the percentage of recombinants were higher, we would know that the loci were further apart on the chromosome. The term locus (plural, loci) is used to describe the position of a gene on a chromosome.
AN EXAMPLE OF PHENOTYPIC RATIO THAT INDICATES LINKAGE
| Phenotype | Approx. Expected numbers if no linkage (9:3:3:1) | Observed numbers in F2 generation |
| Purple, elongated | 405 | 336 |
| Purple, rounded | 135 | 31 |
| Red, elongated | 135 | 28 |
| Red, rounded | 45 | 325 |
| Total | 720 | 720 |
When genes are not linked, we expect them to
be separated 50 per cent of the time because there is a 50 per cent chance that
alleles on separate chromosomes will be inherited together. When looking for
linkage, geneticists look for a frequency of recombinants significantly less than
50 per cent.
CHROMOSOME MAPS
The frequency with which linked genes are separated by crossover can tell us a lot about the relative positions, or loci, of these genes on the chromosome. Two alleles that are close together are rarely separated by crossing over, while those located on opposite ends of the chromosome are separated almost every time. Analysis of crossover frequency can be used to work out the relative positions of alleles and so make chromosome maps.
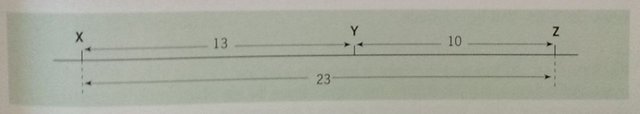
An image drawn by me
Genes Y and Z have a crossover frequency of 10 per cent and so are 10 units apart. A third cross between Y and X determines the relative positions of X, Y and Z. If the order of genes is XYZ (as shown), the crossover frequency between X and Y is 13 per cent. If the order is XZY, the crossosver frequency is 33 per cent.
SEX DETERMINATION
What decides whether a baby will be a boy or a girl?
In humans, sex determination depends on the inheritance of special sex chromosomes. Every human cell contains 23 pairs of chromosomes: 22 pairs of autosomes and one pair of sex chromosomes. Females have two large X chromosomes and so we say they are the homogametic sex. Males have one X chromosome and one smaller Y chromosome, and are therefore the heterogametic sex.
In the female, all eggs receive an X chromosome during meiosis. However, meiosis in the male produces sperm, half of which receive a chromosome and half an X. So as the sperm swim for the female’s egg it’s a real race: if a Y sperm fertilizes the female’s egg, the offspring is male. If an X sperm wins, the baby will be a female.
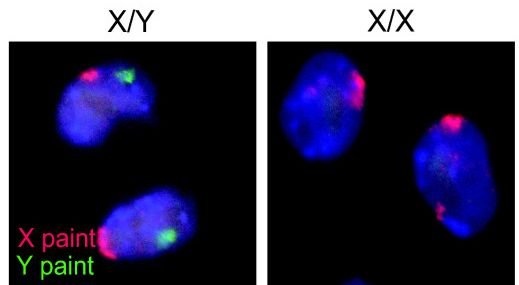
In humans, about 114 boys are conceived for every 100 girls. We do not really understand the reason for this. Interestingly, however, more male embryos die and so at birth the proportions are down to 106 to 100. By puberty the numbers are equal and in old age the females outnumber males by two to one.
In the 1990s, a group of genetic researchers demonstrated that the Y chromosome carries just 60 functional genes, including a male-determining gene called SRY. All embryos are female unless the active SRY imposes maleness upon it. When a male embryo and a female embryo share the same blood supply, it is possible that the SRY can produce hormones that can force an XX embryo to develop as a male. This is an extremely rare situation in humans but it is common in cattle.
Androgen insensitivity syndrome (AIS) is condition that can cause a genetically male child to develop outwardly as a female. The receptor proteins for the male hormones (androgens) don’t work properly, so the embryo fails to read the signals that should turn it into a male. This syndrome may go undetected until the teens, when puberty fails to happen. Genetic analysis will reveal that the ‘girl’ is in fact a male. Although she will look and act like a girl, she will not menstruate and will be unable to have children.
SEX-LINKED INHERITANCE
Genes carried on the sex chromosomes are said to be sex-linked. Human females have two X chromosomes, which, like the autosomes, carry two alleles of every gene. Females therefore have two sets of sex-linked alleles. In males, however, the Y chromosome is smaller and cannot mirror all of the genes on the X chromosome, so males have only one set of most sex-linked alleles. This is why males suffer from the effects of X-linked genetic diseases more often than females. There are no known Y-linked diseases or conditions, probably because the Y chromosome carries so few genes.
A classic example of a sex-linked trait transmitted by the X chromosome is haemophilia. People suffering from this disease do not make factor VIII, an essential component in the complex chain reaction of blood clotting. In addition to the problems caused if they injure themselves, haemophiliacs suffer from internal bleeding as a result of normal activity. Bleeding at the joints during even light exercise is a particular problem. However, haemophiliacs can usually live a full and active life by having regular injections of Factor VIII.
Haemophilia is caused by a sex-linked recessive allele. Females have a pair of alleles but males possess only one. So, if a male inherits the haemophilia allele, he has the disease since he cannot possess another healthy allele to mask its effect.
Using the following notation:
Xh = the haemophilia allele on the X chromosome
XH = the healthy allele on the X chromosome
Y = the Y chromosome which does not carry either allele
We can see that the possible genotypes are:
XHXH = healthy female
XHXh = healthy, carrier female
XHY = healthy male
XhY = haemophiliac male
The inheritance of haemophilia can be studied using a pedigree diagram. The word pedigree means ‘of known ancestry’ and applies as much to humans as it does to domestic animals.
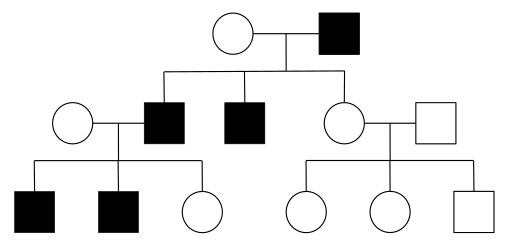
The pedigree of the Royal Family can be traced back to 1066 and shows that Queen Victoria was a carrier of the haemophilia allele and passed it on to four of her nine children. Other sex-linked traits in humans include Duchenne muscular dystrophy (DMD) and red-green colour blindness. A person with red-green colour blindness cannot distinguish between red, green, orange, and yellow. On this Ishihara test charts, a person with normal vision would see numbers but a person with red-green colour blindness would see only a random pattern of dots. This condition is caused by a defective allele that codes for the one of the three groups of light-sensitive cone cells in the retina. Eight per cent of males but only 0.4 per cent of females suffer from colour blindness.
Thanks for reading.
REFERENCES
https://en.wikipedia.org/wiki/Cystic_fibrosis
https://www.mayoclinic.org/diseases-conditions/cystic-fibrosis/symptoms-causes/syc-20353700
https://www.medicalnewstoday.com/articles/147960.php
http://www.nature.com/scitable/topicpage/gregor-mendel-and-the-principles-of-inheritance-593
https://study.com/academy/practice/quiz-worksheet-characteristics-of-a-dihybrid-cross.html
https://www.sanfoundry.com/cytogenetics-questions-answers-dihybrid-cross/
https://www.slideshare.net/HiwrHastear/genetic-linkage-and-crossing-over
https://courses.lumenlearning.com/boundless-biology/chapter/laws-of-inheritance/
https://www.biology.iupui.edu/biocourses/N100/ch3mendelexcept.html
https://en.wikipedia.org/wiki/Locus_(genetics)
https://examples.yourdictionary.com/examples-of-genotype-phenotype.html
https://en.wikipedia.org/wiki/Genetic_linkage
https://www.k-state.edu/parasitology/biology198/answers2.html
http://www.nature.com/scitable/topicpage/epistasis-gene-interaction-and-phenotype-effects-460
@tipu curate
Posted using Partiko Android
Upvoted 👌 (Mana: 5/20 - need recharge?)
thanks.
Hi, I'm riccc96, a curator of stem.curate, I found your post and I had the pleasure to read it and appreciate it, you got our 100% vote if you like to come and visit us we are happy to welcome you with open arms
riccc96
Hello,
Your post has been manually curated by a @stem.curate curator.
We are dedicated to supporting great content, like yours on the STEMGeeks tribe.
If you like what we are doing, please show your support as well by following our Steem Auto curation trail.
Please join us on discord.
This post has been voted on by the SteemSTEM curation team and voting trail. It is elligible for support from @curie and @minnowbooster.
If you appreciate the work we are doing, then consider supporting our witness @stem.witness. Additional witness support to the curie witness would be appreciated as well.
For additional information please join us on the SteemSTEM discord and to get to know the rest of the community!
Thanks for having used the steemstem.io app and included @steemstem in the list of beneficiaries of this post. This granted you a stronger support from SteemSTEM.