My shittiest method/experiment: Yeast Transformation

@sco is doing a "my shittiest method" challenge thing and I have been nominated by @scienceangel ! Took me some time but now I am writing it. Although it's more of a "my shittiest experiment" than "my shittiest method", as the method was technically fine.
The rules for this
- describe the worst method/assay/experiment you have to work
- use the #myshittiestmethod and the #steemstem tag
- use “My shittiest assay method: … ” as title
- you can nominate some peers, they have to participate then (unless they want to be seen as very, very lame). But you can join the party without invitation as well!
So, here we go.
Some Background
At the time, I had a 7 weeks internship in a microbiology lab (about which I wrote one of my first posts, the only post of mine ever to be curied!), where they were researching specific protein channels in the membrane of the ER.
What?
The ER (or Endoplasmic Reticulum) is a large part of eukaryotic cells - cells that have a nucleus (so no bacteria!). It has several functions, making sure proteins are folded correctly is one of them because proteins usually need a 3-dimensional structure to function as intended.
On the membrane of the ER, there are several channels that are responsible for transporting proteins into and out of the ER, because simple diffusion would be way too slow. In yeast, one of these channels is called SEC61.
The question in my experiment was, if the overexpression (= "cell uses a specific gene more than usual to produce a specific protein more than usual") of specific proteins has an effect on the export out of the ER when certain protein channels don't work anymore.@suesa
Why?
Well, nobody is 100% sure how much SEC61 and other channels are involved in said export.
But why do we care?
In humans, a similar channel in the ER-membrane exists. Proteins that are wrongly folded in the ER (it's a biological system, there are always mistakes) need to be transported to the outside and taken apart again, or the cell might die.
There are a lot of diseases, especially neurodegenerative diseases like Alzheimer's disease are linked to the aggregation of misfolded proteins. Knowing how the cell transports them and how to support or fix this system is thus something very interesting.
And it's easier to test the basics in yeast, before doing weird stuff to humans, you know?
The Experiment
The insertion of the genes to overexpress the proteins we wanted to check into the "damaged" yeast strains was a piece of cake and worked without any further issues.
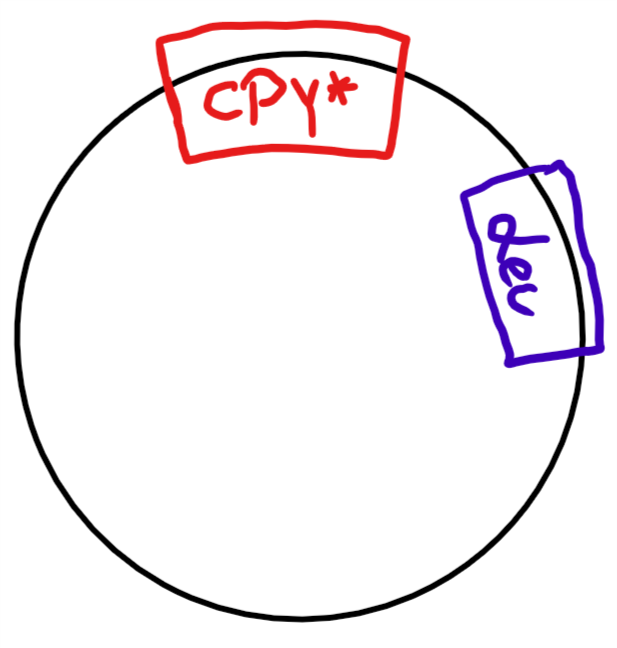
The real problem was CPY*. CPY* is the mutated version of a gene that usually codes for a protein destined for the secretory pathway. As such, it's perfect to monitor how a cell handles misfolded proteins.
In theory, you create a plasmid (round piece of DNA), which includes the gene you want (CPY*) and a so-called marker gene. A marker gene is supposed to tell you if the organism you're transforming (=inserting DNA into) absorbed the DNA or not. It can be almost anything, bacteria are usually transformed with a marker gene for antibiotic resistance.
For the yeast, we used a gene that allowed the organism to produce Leucine (an amino acid) by itself, thus enabling it to grow in a petri dish with nutritional agar missing Leucine (= Leucine dropout medium).
My first step was to take some E. coli, into which the plasmid had been transformed. E. coli is great for multiplying Plasmids, so you don't have to build a thousand of them by hand.
The E. coli grow overnight in a liquid medium, then the plasmids are isolated by destroying the cells (the process took about 45 minutes after I got used to doing it). Then, the purified plasmids are transformed into the yeast (1.5 to 2 hours, depending on how good my day was going) and the yeast can grow for 2-3 days on the dropout medium.
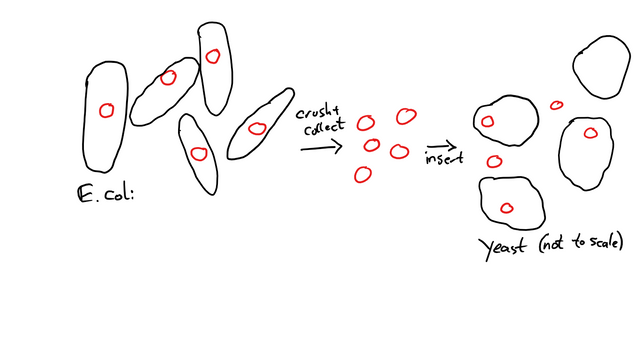
As you can see, not every yeast cell absorbed a plasmid during this process. That's why we need the marker gene and the dropout medium: We only want yeast cells that have been successfully transformed.
In theory, my plate with the yeast was supposed to look like this after 3 days:
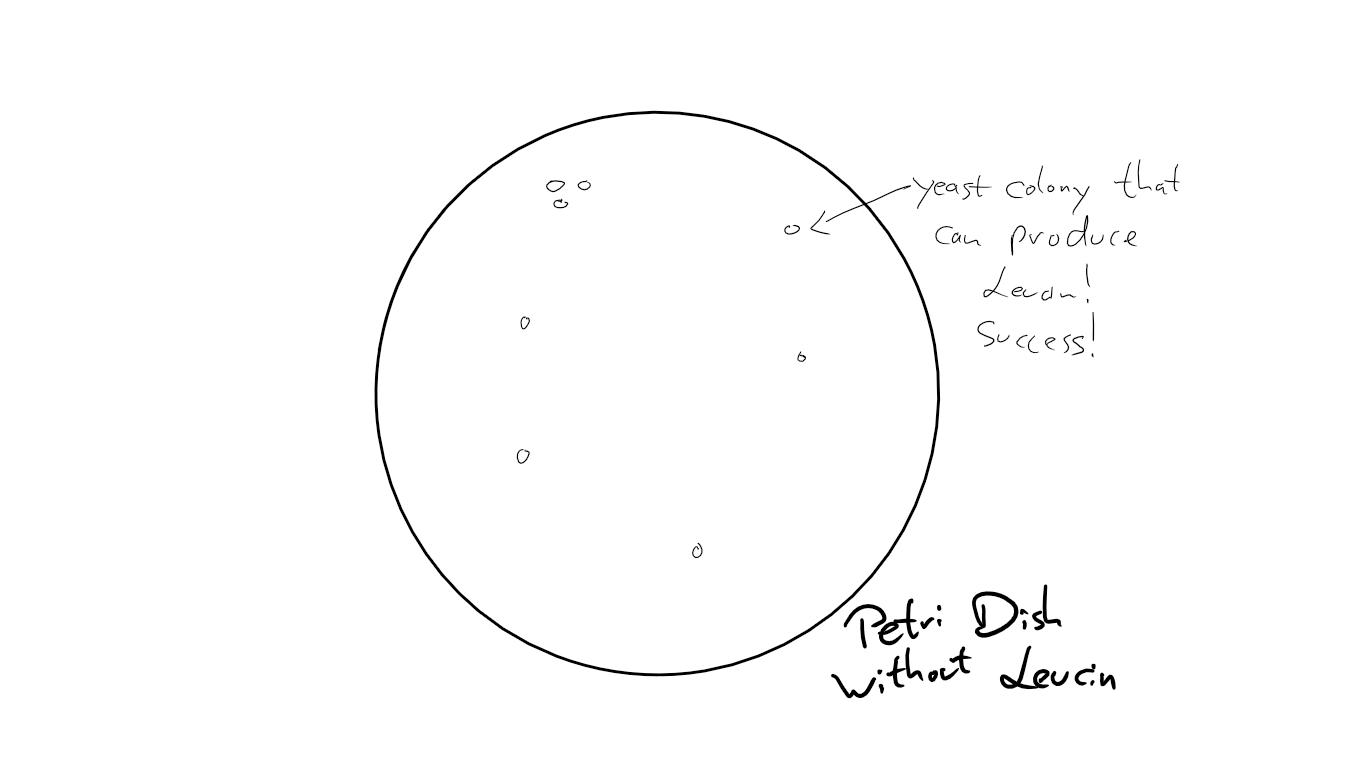

The sad truth was ... nothing happened.
But I was a 4th-semester student with not a lot of lab experience, I had probably just done something wrong! Maybe the yeast hadn't properly absorbed the plasmid, after all, it was a foreign gene and yeast isn't so super happy about introducing foreign DNA into its cell.
I repeated the process.
And then I repeated it again.
And again.
At this point, my supervisor did the extraction of the plasmid and yeast transformation himself, to give me the chance to continue with my research project.
He failed too.
That was the point when we considered something might be off. 4 weeks had already passed of the 7 I had been given in that lab and we weren't even close to getting results!
To find out what might be wrong, we did a PCR to check if the gene had been successfully inserted.
This means we took some of the yeast cells and duplicated their DNA by heating it so it separates into two strands, which then were completed to new DNA strands, effectively doubling the amount of available Gene DNA.
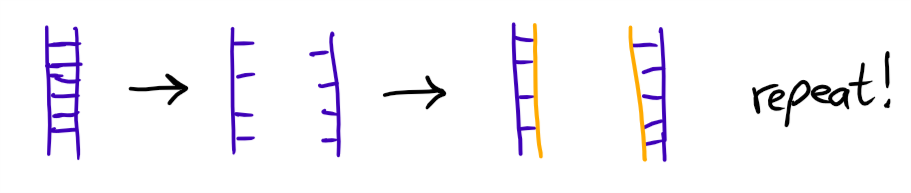
That's done several times (the whole thing took about 2 hours), then the DNA is put on a gel for a gel electrophoresis (please just read the Wikipedia entry).
The CPY* gene was there.
So we tried again.
And failed. Both.
So we repeated the PCR and sent some samples to a company, to have them sequence the plasmid and tell us if something is off.
The CPY* gene was there.
Our transformation didn't work.
I wonder if one of you guys already has an idea what might have gone wrong? We only noticed our mistake in week 6.
All the time, we had been checking if the CPY* gene was there. We never checked for the marker gene, because why should we? That's supposed to be pretty stable.
We did a PCR to check for the marker gene. It was not there. The person who had built this construct (another student, apparently) had managed to build a plasmid without marker gene and thus ruined 7 weeks of work for me.

That day, I sat at the computer in the lab, looking at the results and just thinking ...

I learned two things during these 7 weeks:
- When you're troubleshooting, check all possible ways something might have gone wrong, not just the most likely
- Do everything yourself. Trust no one. I could have built that plasmid myself, transformed the yeast AND done my testing in those 7 weeks. Instead, I was left with no results and slight disappointment.
Conclusion
As I said at the beginning, it's not really my shittiest "method", but most of my time in the lab has been spent with experiments tailored to teach students something, not to annoy them. I am sure there are a lot more shitty methods to come
As the next person, I am nominating @ertwro !
Pictures by me, except for the first one, that's from pixabay.com

At first i was lost then i got the last part. I work on cars for a living and i have learned the hard way to trust no one. I have to pry out of customers what they have screwed up in thier missguided attempts to fix what the actual problem was in the first place. My moto is
It's not just that everybody lies (they do) but everybody fucks up! :D
That is true, I try to be perfect in my job everyday but sometimes i screw up and then i beat myself up over it. One of my old bosses told me once " if you never screw up i would have to fire you because you are obviously not human".
Hahaha the best costumers that bring me their broken bikes are: I need this and this fixed. I already started repairing it to "help you a little bit" (and messing everything up and not telling you)
I lost so many hours because of these people..
we call those " basket cases " because they bring everything in, in a basket and expect you to do it for half price since they already took it apart.

LOL
really grinds my gears... Once a costumer cound not fix his motorcycle and came to me. I asked him what he did, how it happended and what he tried to do to fix it. Did not tell me much and I nearly disassembled the whole electronic until I found after many hours of work, that he installed the Inclination sensor upside down.... could kill him for that. But he paid every single hour for the troubleshooting
oh yeah, that kind of stuff is the absolute worst. When people get into their fuse panels is always an adventure to. i have seen so many people get their asses kicked by someone pulling a fuse and putting it back in the wrong spot. GRRRR
just normal for mechanics. I just repair like 4 bikes a month, but you have do do way more. I would kick so many asses if I were you ahahaha
PS: House is the best!!!
I love house. that show is pure gold.
unfortunally there will be no more episodes because hugh laurie found it too exhausting... daim... it was so goooood and he has the same taste of sarcasm as I have.
miss it so hard
YES! the sarcasm is what made it.
Aahhahahahahahahaha
I mean, this truly sucks. They sometimes say people should work at least once through the entire process so they can see the effects what will happen if they fuck something up.
Now you lost too many weeks because someone fucked up and they might never know they messed up something important.
Life is rought.
Lol, classic. A well-established method doesn't work when applied by a student... You roll your eyes...
damn incompetent students, I've got better stuff to do
"Repeat it, please!"
...
"What, it didn't work again?"
Wow, that one really IS incompetent...but he/she looked so nice at first...strange...I'll show them how it works
You do it yourself and it doesn't work.
Whaaaaaat??!?
Happens like every single time.
And that's where your supervisor made the error. Never trust your student. Not even a good one. Those supersticions are there for a reason ;-)
Thanks for participating! Even if I fear it won't be your worst experiment by end-PhD-time, it was for sure a pretty disappointing one - but one that thought you a very important lesson about science too! Don't blindly trust others to work well.
Tbh, it was a fun time. I'm now super good at yeast transformation, working sterile and doing the same damn thing every day.
Haha, that's kind of the story of my research life.
Well, you did learn something!
Just not anything you were looking to learn.
@suesa : I loved this post very much.
So sad. Working with a group of people is good and bad. Sometimes it gives your great success, sometimes you struggle. I can understand how difficult it would be in a wet lab compared to a computational lab. By the way I thought you are an undergrad. (Or Am I wrong?) To get these experiences in the early stage of your career is great.
Yes, I don't have any degree yet :) but the labs at our university are generally willing to take in students for a few weeks, if they ask ahead of time. Most just never take the chance because there's no time. I was lucky that I had somr extra time to do this!
Atleast you learned something 😂
ER about it I remember it got ribosomes and they help in protein synthesis and ya it's rough n smooth .. Good luck with your exp
The fact that you guys tried and fail didn't make this experiment shitty in itself because experience matters, more so when the fault wasn't directly yours.
This challenge on its own is quite commendable as it recognizes the fact that science is not free of errors; the question: describe the worst experiment...
Interesting experience
@eurogee of @euronation community & The Host, Show Us Your Witnesses Weekly Contest
In this life dont trust no one !! Better 🤣