The Citizen Science Grid projects #1 - DNA@home. BOINC volunteer computing.
The Citizen Science Grid is a project, which is initiated by Travis Desell, an Assistant Professor in the Computer Science Department, University of North Dakota. In a series of posts I will represent the project's details or its sub-projects particularly.
The Citizen Science Grid "hosts" 4 different projects
- DNA@home
- SubsetSum@Home
- Climate Tweets
- Wildlife@Home
Do you want to study DNA right at your home?
DNA@home
You cannot really make studies about DNA (or maybe you can, if you a biologist, doctor or sth.), but your computer can make! Of course, if you join the Citizen Science Grid project by using the BOINC software, developed by the University of Berkeley.
DNA@Home is a collaboration between the Computer Science Department and the Basic Sciences Department of the University of North Dakota. It has been also developed with a lot of help from Rensselaer Polytechnic Institute. The project's goal is to discover the regulations of the the genes in DNA and what exactly regulates them. You know of course, that skin cells are different from the muscle ones, or bone ones, even though all these cells have every gene in your genome? All the genes play together all at once. Depending on what cells are trying to do at any particular time, only a unique part of the genes are used, and the remainder are somewhat isolated, or turned off. DNA@home uses statistical algorithms to unlock the key to this complex regulations, using your volunteered computers.
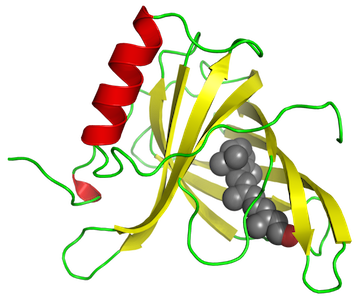
The way of regulating genes at the stage of "transcription" where a molecule called a polymerase makes "reading" along DNA from the start to the end of the gene and creates an RNA, which is a type of "messenger". Other molecules - transcription factors, join to the DNA at the beginning of the gene and potentially help to recruit the polymerase. It is the presence or absence of the binding of these transcription factors that determine whether a gene is shut down, or must work! That's where scientists do not know exactly which are the transcription factors responsible for regulating particular genes.
Transcription factors use so-called "fingers" in order to find a certain short pattern in the nucleotides , which represent the letters the sequence of DNA, but in many cases scientists don't know what exactly are the patterns. The software for the whitelisted project searches some short sequences of nucleotides looking more-or-less the same near multiple gene beginnings. They also appear comparatively the same in some corresponding places in the genomes of species, which are related in a way. The problem is, that DNA sequences are enormous and ranging from millions to billions of nucleotides. The sequences are short and only approximately conserved from one site to the nextwhich is almost an impossible task if we don't share a lot of computation power with the project.
This is where all our help and computer resources will be very valuable, and we should consider giving this project a boost. Don't just waste the resources and computation power in pointless mining of various coins. One more thing! Getting involved in BOINC and Gridcoin whitelisted projects will show some profit. The generous network will reward you with precious Gridcoin.
So what a better time to dive into the Science and Medicine pool, generously delivered by BOINC and Gridcoin?
Don't waste more time! See you with the next project update next week. Bye!
Sources:
https://www.gridcoin.us/
https://csgrid.org/csg/ - images and valuable information
https://boinc.berkeley.edu/ - easily download and install the BOINC software

Thank you for the breakdown.I'll check out how much magnitude I can get with this and I might switch from my current project if it's similar.
You just planted 0.09 tree(s)!
Thanks to @ragnarokdel
We have planted already 3364.58 trees
out of 1,000,000
Let's save and restore Abongphen Highland Forest
in Cameroonian village Kedjom-Keku!
Plant trees with @treeplanter and get paid for it!
My Steem Power = 18543.42
Thanks a lot!
@martin.mikes coordinator of @kedjom-keku
=======================================================================================
This post was upvoted by Steemgridcoin with the aim of promoting discussions surrounding Gridcoin and science.
This service is free. You can learn more on how to help here.
Have a nice day. :)